White light CCF of HD 189733 - two nights
In this notebook, we present a simple example of the application of the Doppler Shadow technique to two nights of ESPRESSO observations of HD 189733, a K1V star, extracting the white light CCFs provided by the DRS. The nights of observations were 11-08-2021 and 31-08-2021.
We run two different instances of HECATE and then use the multi_night_analysis class to combine the data.
[1]:
import numpy as np
from HECATE.HECATE import HECATE # main class
from HECATE.get_data import *
from HECATE.multi_night_analysis import multi_night_analysis # to perform the multi-night analysis
Warming up JIT-compiled functions...
[2]:
stellar_params = {
"Teff":4969, "Teff_err":43, #effective temperature [K]
"logg":4.60, "logg_err":0.01, #superficial gravity [dex]
"FeH":-0.07, "FeH_err":0.02, #metallicity [dex]
"P_rot":2.21857312, #rotation period [d]
"R_star":0.766, #radius [solar radii]
"inc_star":71.87 #stellar inclination [º]
}
planet_params = {
"P_orb":2.21857312, #orbital period [d]
"a_R":8.76863, #system scale [stellar radii]
"Rp_Rs":0.1602, #planet-to-star radius ratio
"t0":53988.30339, #mid-transit time [d]
"e":0, #orbital eccentricity
"w":90, #argument of periastron [º]
"inc_planet":85.465, #planet inclination [º]
"lbda":-1.00, #spin-orbit angle [º]
"dfp": -0.002424 #mid-transit phase shift, night of 2021-09-11
}
Now we quickly run HECATE for the first night of observations, saving the local CCF parameters.
[3]:
CCFs, time, airmass, berv, bervmax, snr, list_ccfs = get_CCFs(planet_params, stellar_params,
day='2021-08-11',
directory_path="HD189733_ESPRESSO_white_light_ccfs",
plot=False)
[4]:
hecate11 = HECATE(planet_params, stellar_params, time, CCFs, spectra=None, plot_soap=False)
[5]:
plot = {"fits_initial_CCF":False, # plot the initial fits to the CCFs
"sys_vel_ccf":False, # plot the systemic velocity of the star in the CCFs
"avg_out_of_transit_CCF":False, # plot the average out-of-transit CCF
"local_CCFs":True, # plot the local CCFs
"photometrical_rescale":False # if the local CCFs are rescaled by the photometric transit model
}
ccf_type = "white light"
model_fit = "modified Gaussian"
[6]:
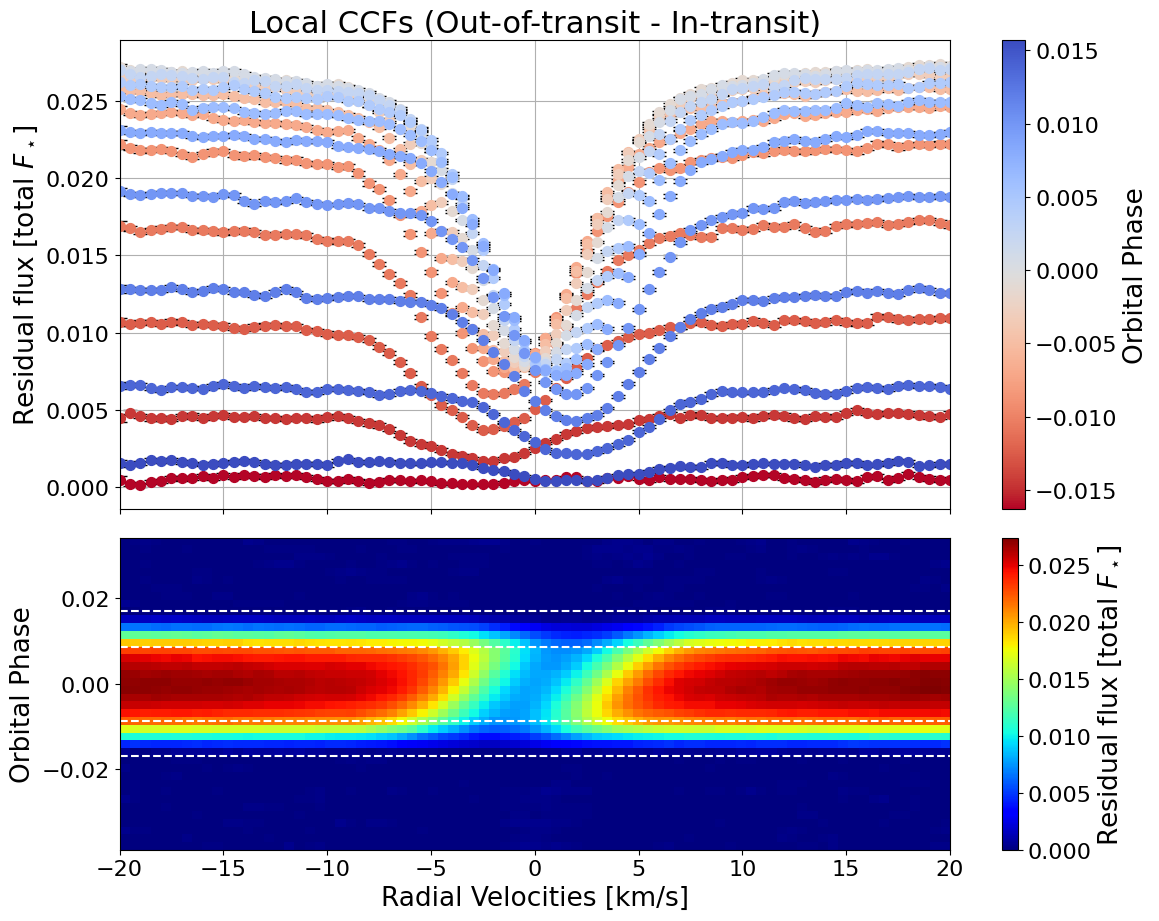
local_CCFs, CCFs_flux_corr, CCFs_sub_all, avg_out_of_transit_CCF = hecate11.extract_local_CCF(model_fit, plot, save=None)
[7]:
master_results11 = hecate11.get_profile_parameters(profiles=avg_out_of_transit_CCF,
data_type="CCF",
observation_type="master",
model="modified Gaussian",
print_output=False,
plot_fit=False)
[8]:
local_results11 = hecate11.get_profile_parameters(profiles=local_CCFs,
data_type="CCF",
observation_type="local",
model="modified Gaussian",
print_output=False,
plot_fit=False)
[9]:
indices_final11 = np.array(list(set(np.where(local_results11['R2'] >= 0.95)[0]).intersection(set(np.where(hecate11.mu_in >= 0.3)[0]))))
local_params11 = [local_results11['central_rv'], local_results11['width'], local_results11['intensity']]
master_params11 = [master_results11['central_rv'], master_results11['width'], master_results11['intensity']]
Now we do the same for the second night.
[10]:
planet_params["dfp"] = -0.002300 # mid-transit phase shift, night of 2021-08-31
[11]:
CCFs, time, airmass, berv, bervmax, snr, list_ccfs = get_CCFs(planet_params, stellar_params,
day='2021-08-31',
directory_path="HD189733_ESPRESSO_white_light_ccfs",
plot=False)
[12]:
hecate31 = HECATE(planet_params, stellar_params, time, CCFs, None)
plot = {"fits_initial_CCF":False,
"sys_vel_ccf":False,
"avg_out_of_transit_CCF":False,
"local_CCFs":True,
"photometrical_rescale":False}
ccf_type = "white light"
model_fit = "modified Gaussian"
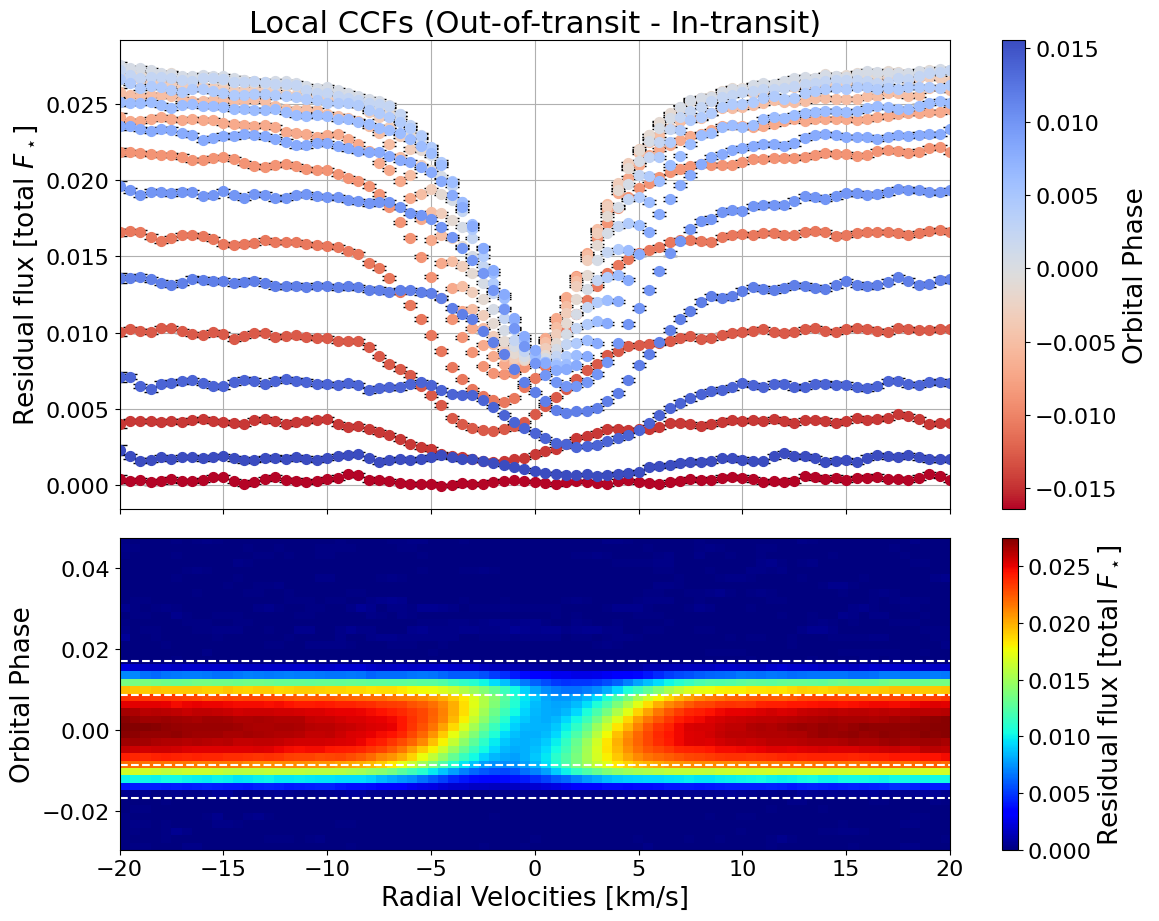
local_CCFs, CCFs_flux_corr, CCFs_sub_all, avg_out_of_transit_CCF = hecate31.extract_local_CCF(model_fit, plot, save=None)
[13]:
master_results31 = hecate31.get_profile_parameters(profiles=avg_out_of_transit_CCF, data_type="CCF",
observation_type="master", model="modified Gaussian",
print_output=False, plot_fit=False)
local_results31 = hecate31.get_profile_parameters(profiles=local_CCFs, data_type="CCF",
observation_type="local", model="modified Gaussian",
print_output=False, plot_fit=False)
[14]:
indices_final31 = np.array(list(set(np.where(local_results31['R2'] >= 0.9)[0]).intersection(set(np.where(hecate31.mu_in >= 0.3)[0]))))
local_params31 = [local_results31['central_rv'], local_results31['width'], local_results31['intensity']]
master_params31 = [master_results31['central_rv'], master_results31['width'], master_results31['intensity']]
Having the results for each night, we build the following dictionaries:
[15]:
night_data_11 = {
'hecate': hecate11, # HECATE instance
'indices': indices_final11, # indices of points to retain
'local_params': np.array(local_params11), # local CCF parameters
'master_params': np.array(master_params11), # master CCF parameters
'color': "blue", # color to use in the plots for this night
'label': "2021-08-11" # label to use in the plots for this night
}
night_data_31 = {
'hecate': hecate31,
'indices': indices_final31,
'local_params': np.array(local_params31),
'master_params': np.array(master_params31),
'color': "red",
'label': "2021-08-31"
}
[16]:
night_data = {"2021-08-11": night_data_11,
"2021-08-31": night_data_31}
[17]:
mult_night = multi_night_analysis(night_data, data_type='CCF')
We can just plot the two nights data in one single plot for each position parameter (\(\phi\) or \(\mu\)):
[18]:
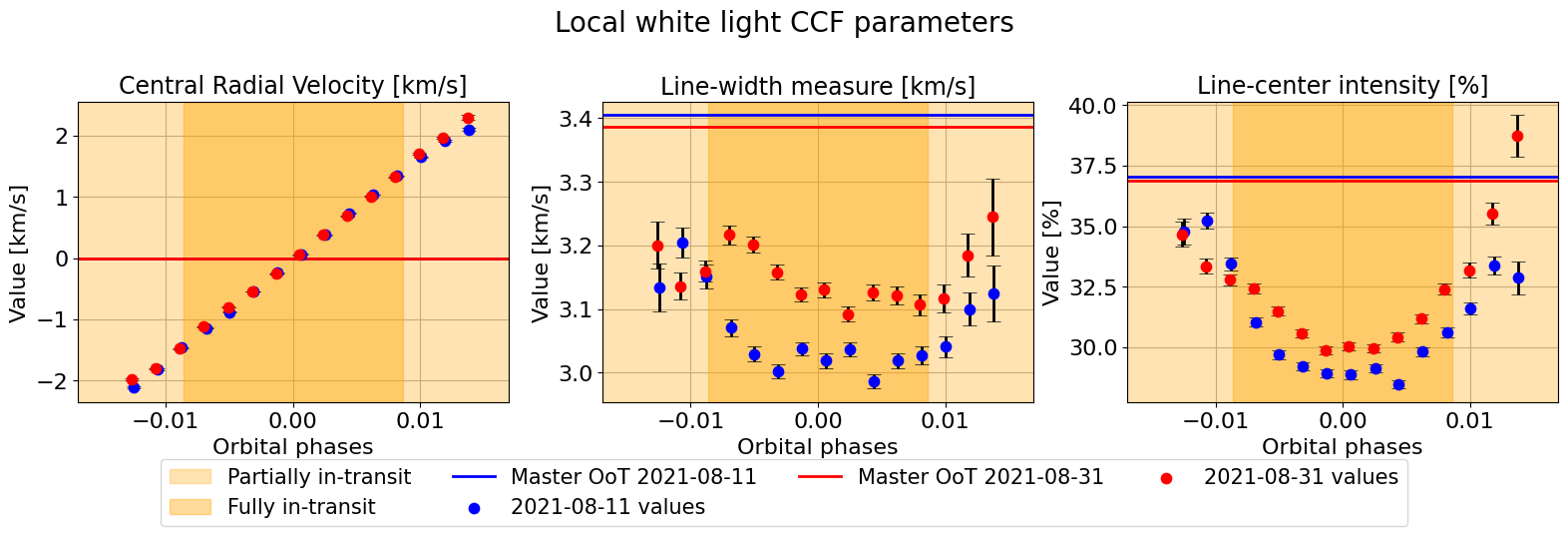
mult_night.plot_parameters(param_type='phases',
fit_each_night=False,
fit_combined=False,
combined_night_names=None,
plot_nested=False,
suptitle="Local white light CCF parameters")
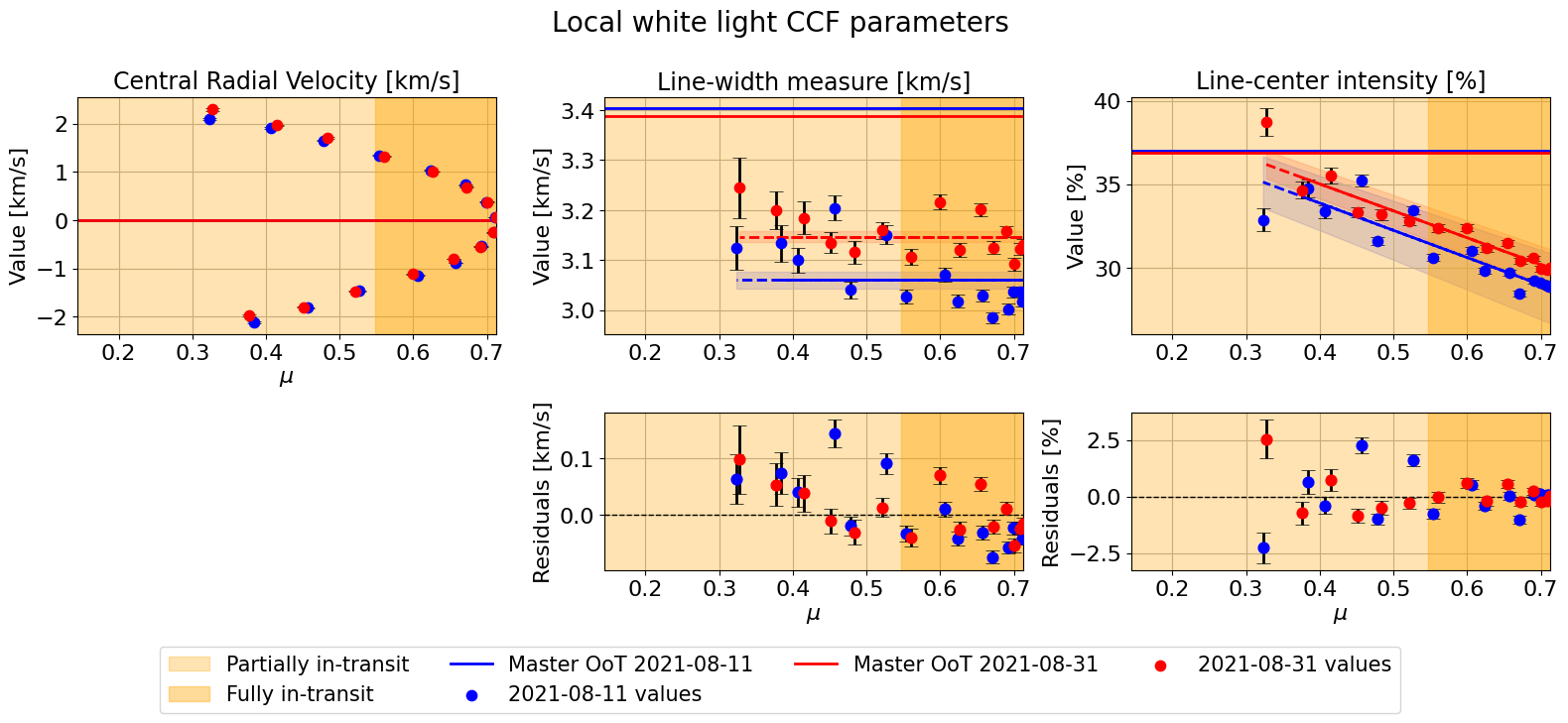
mult_night.plot_parameters(param_type='mu',
fit_each_night=False,
fit_combined=False,
combined_night_names=None,
plot_nested=False,
suptitle="Local white light CCF parameters")
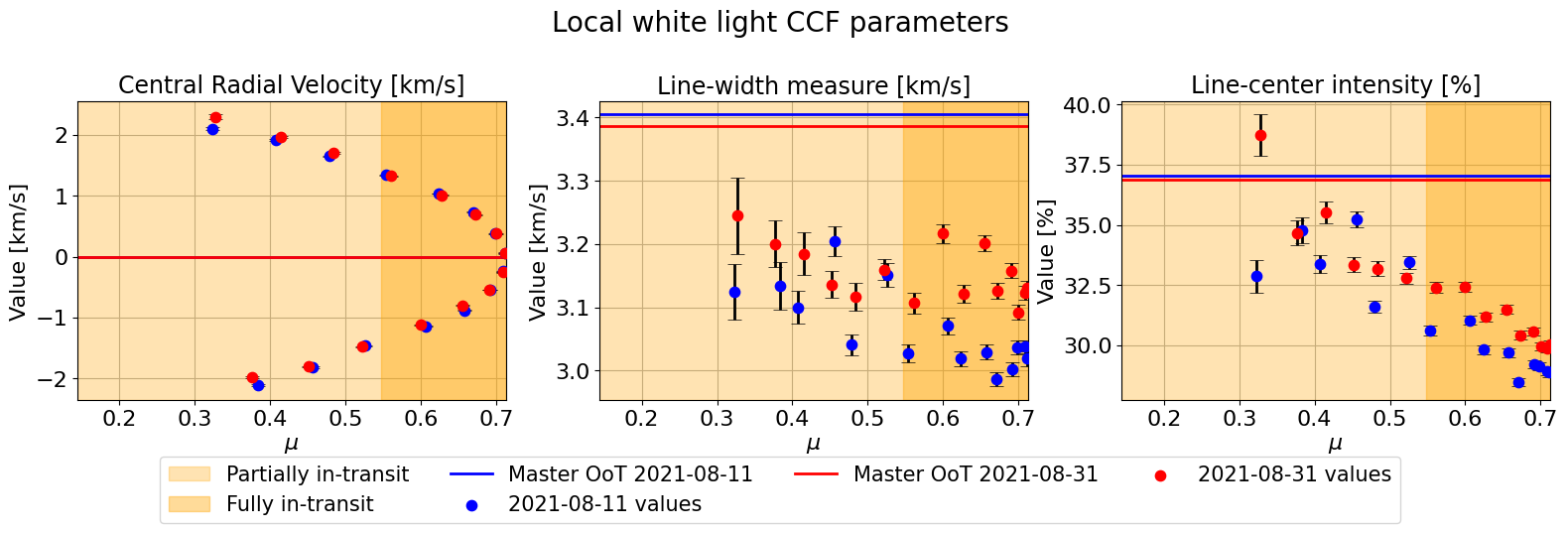
[18]:
{}
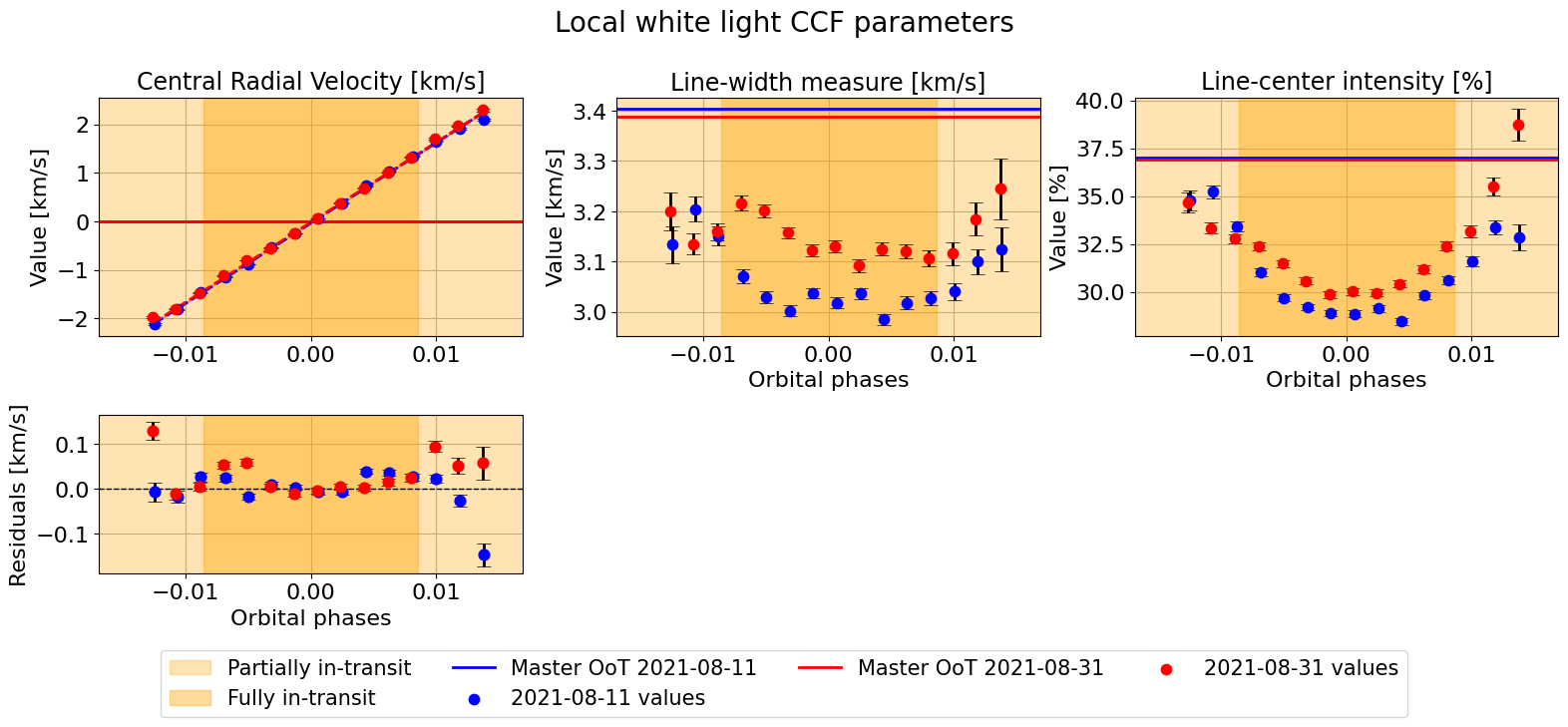
Or fit a linear model to each night:
[19]:
fit_results = mult_night.plot_parameters(param_type='phases',
fit_each_night=True,
fit_combined=False,
fit_param_indices = np.array([0]),
suptitle="Local white light CCF parameters")
21190it [00:31, 663.00it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 442044 | eff(%): 4.677 | loglstar: 41.167 < 46.444 < 45.864 | logz: 26.002 +/- 0.124 | stop: 0.897]
17090it [00:22, 770.34it/s, batch: 7 | bound: 3 | nc: 1 | ncall: 332341 | eff(%): 4.985 | loglstar: -17.931 < -12.568 < -13.830 | logz: -24.281 +/- 0.087 | stop: 0.884]
------------------------------
Linear vs Constant
logZ(linear) = 26.002 ± 0.113
logZ(constant) = -24.257 ± 0.083
log Bayes factor = 50.259
Unconstrained model favored.
------------------------------
21770it [00:33, 652.21it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 455201 | eff(%): 4.670 | loglstar: 41.207 < 46.447 < 45.843 | logz: 24.882 +/- 0.127 | stop: 0.898]
17613it [00:24, 730.22it/s, batch: 5 | bound: 6 | nc: 1 | ncall: 359350 | eff(%): 4.756 | loglstar: -17.751 < -11.014 < -12.864 | logz: -27.839 +/- 0.125 | stop: 0.979]
Positive vs Negative slope
===========================
logZ(m>0) = 24.878 ± 0.116
logZ(m<0) = -27.826 ± 0.109
log Bayes factor = -52.704
Positive slope favored.
==========================
Linear fit parameters:
m = 165.446158 +/- 1.393267
b = -0.037505 +/- 0.004496
ln_f = -3.522776 +/- 0.232864
------------------------------
22094it [00:33, 655.36it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 461484 | eff(%): 4.676 | loglstar: 38.304 < 43.603 < 43.184 | logz: 21.629 +/- 0.129 | stop: 0.890]
16351it [00:21, 758.91it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 316766 | eff(%): 4.996 | loglstar: -17.750 < -12.542 < -13.195 | logz: -24.193 +/- 0.092 | stop: 0.940]
------------------------------
Linear vs Constant
logZ(linear) = 21.645 ± 0.116
logZ(constant) = -24.186 ± 0.084
log Bayes factor = 45.832
Unconstrained model favored.
------------------------------
21821it [00:32, 664.49it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 454749 | eff(%): 4.685 | loglstar: 38.395 < 43.597 < 43.144 | logz: 21.769 +/- 0.130 | stop: 0.901]
19592it [00:27, 719.54it/s, batch: 6 | bound: 7 | nc: 1 | ncall: 405877 | eff(%): 4.700 | loglstar: -18.435 < -10.892 < -12.594 | logz: -27.248 +/- 0.115 | stop: 0.783]
Positive vs Negative slope
===========================
logZ(m>0) = 21.774 ± 0.117
logZ(m<0) = -27.237 ± 0.103
log Bayes factor = -49.011
Positive slope favored.
==========================
Linear fit parameters:
m = 164.633395 +/- 1.720329
b = -0.020113 +/- 0.005087
ln_f = -3.285271 +/- 0.228280
------------------------------
[20]:
fit_results = mult_night.plot_parameters(param_type='mu',
fit_each_night=True,
fit_combined=False,
fit_param_indices = np.array([1, 2]),
suptitle="Local white light CCF parameters")
21858it [00:33, 659.64it/s, batch: 6 | bound: 3 | nc: 1 | ncall: 456539 | eff(%): 4.675 | loglstar: 35.622 < 40.547 < 39.524 | logz: 16.645 +/- 0.137 | stop: 0.980]
18196it [00:26, 694.17it/s, batch: 8 | bound: 3 | nc: 1 | ncall: 357714 | eff(%): 4.942 | loglstar: 30.035 < 34.704 < 34.135 | logz: 21.300 +/- 0.091 | stop: 0.891]
------------------------------
Linear vs Constant
logZ(linear) = 16.623 ± 0.128
logZ(constant) = 21.296 ± 0.086
log Bayes factor = -4.673
Zero-slope model favored
========================
Linear fit parameters:
b = 3.060385 +/- 0.016268
ln_f = -3.952245 +/- 0.217799
------------------------------
21623it [00:33, 649.11it/s, batch: 6 | bound: 3 | nc: 1 | ncall: 451080 | eff(%): 4.679 | loglstar: 36.649 < 41.633 < 40.313 | logz: 18.027 +/- 0.136 | stop: 0.986]
18467it [00:26, 707.35it/s, batch: 8 | bound: 3 | nc: 1 | ncall: 362837 | eff(%): 4.946 | loglstar: 35.916 < 40.539 < 39.992 | logz: 26.156 +/- 0.095 | stop: 0.911]
------------------------------
Linear vs Constant
logZ(linear) = 18.052 ± 0.128
logZ(constant) = 26.165 ± 0.090
log Bayes factor = -8.112
Zero-slope model favored
========================
Linear fit parameters:
b = 3.146783 +/- 0.010599
ln_f = -4.469513 +/- 0.243132
------------------------------
18897it [00:28, 667.12it/s, batch: 6 | bound: 3 | nc: 1 | ncall: 388140 | eff(%): 4.734 | loglstar: -11.811 < -6.828 < -7.786 | logz: -24.436 +/- 0.116 | stop: 0.961]
16923it [00:23, 715.63it/s, batch: 8 | bound: 3 | nc: 1 | ncall: 330222 | eff(%): 4.967 | loglstar: -24.259 < -19.277 < -19.580 | logz: -29.152 +/- 0.076 | stop: 0.875]
------------------------------
Linear vs Constant
logZ(linear) = -24.450 ± 0.108
logZ(constant) = -29.140 ± 0.071
log Bayes factor = 4.690
Unconstrained model favored.
------------------------------
19234it [00:26, 730.64it/s, batch: 6 | bound: 3 | nc: 1 | ncall: 397422 | eff(%): 4.709 | loglstar: -25.259 < -19.422 < -20.906 | logz: -36.427 +/- 0.114 | stop: 0.876]
18231it [00:27, 673.87it/s, batch: 6 | bound: 4 | nc: 1 | ncall: 373274 | eff(%): 4.744 | loglstar: -11.935 < -6.820 < -7.220 | logz: -23.308 +/- 0.118 | stop: 0.981]
Positive vs Negative slope
===========================
logZ(m>0) = -36.410 ± 0.107
logZ(m<0) = -23.287 ± 0.105
log Bayes factor = 13.123
Negative slope favored.
==========================
Linear fit parameters:
m = -16.288863 +/- 2.245907
b = 40.390411 +/- 1.363256
ln_f = -3.459207 +/- 0.221864
------------------------------
20274it [00:30, 657.75it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 420881 | eff(%): 4.694 | loglstar: -3.788 < 1.518 < 0.887 | logz: -17.367 +/- 0.119 | stop: 0.903]
16225it [00:22, 723.16it/s, batch: 8 | bound: 3 | nc: 1 | ncall: 314015 | eff(%): 5.000 | loglstar: -24.872 < -20.066 < -20.696 | logz: -29.942 +/- 0.078 | stop: 0.911]
------------------------------
Linear vs Constant
logZ(linear) = -17.360 ± 0.110
logZ(constant) = -29.949 ± 0.074
log Bayes factor = 12.588
Unconstrained model favored.
------------------------------
20850it [00:28, 721.94it/s, batch: 7 | bound: 5 | nc: 1 | ncall: 434071 | eff(%): 4.685 | loglstar: -26.534 < -20.084 < -20.927 | logz: -37.472 +/- 0.112 | stop: 0.791]
19908it [00:30, 657.97it/s, batch: 7 | bound: 4 | nc: 1 | ncall: 412332 | eff(%): 4.703 | loglstar: -3.732 < 1.525 < 0.839 | logz: -16.525 +/- 0.116 | stop: 0.901]
Positive vs Negative slope
===========================
logZ(m>0) = -37.467 ± 0.102
logZ(m<0) = -16.539 ± 0.106
log Bayes factor = 20.929
Negative slope favored.
==========================
Linear fit parameters:
m = -16.079402 +/- 1.190150
b = 41.449304 +/- 0.737102
ln_f = -4.388216 +/- 0.347686
------------------------------